|
IGP Organization
The IGP is a collaborative effort between the tick research community, the Microbial Sequencing Centers (MSCs) appointed for the project (the Broad Institute and JCVI), and the NIH. The research community has played an important role in the development of the IGP and community analysis of IGP data is essential. Like all genome projects, the IGP is a continually evolving research project.
The IGP was structured in two phases (I and II) that were conducted simultaneously. Figure I. shows a schematic representation of the organization of the IGP, and a brief description of phase I and II is provided below.
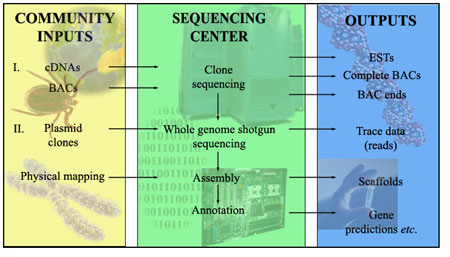
Figure 1. The organization of the Ixodes scapularis Genome Project. Phase I and II activities conducted by the tick research community and the MSCs are shown.
Phase I
Consortium scientists generated many of the tools and resources necessary for the IGP. These included cDNA, Bacterial Artificial Chromosome (BAC) and genomic libraries containing DNA inserts of various sizes. The Broad Institute conducted deep sequencing of two I. scapularis cDNA libraries (library G893: pooled, normalized, non-infected and library G894: pooled, standard, non-infected). These ESTs were used to produce automated gene models (gene predictions) in Phase II. The MSCs jointly shotgun sequenced and assembled 45 BAC clones. These provided an insight into the overall organization of the Ixodes genome, such as the level of heterogeneity in the tick colony selected for the project, % AT content, the type and number of repetitive elements present, and average gene density. The ends of 367,000 BAC clones were also sequenced by the JCVI, providing “sequence-tagged sites” used to help assemble raw genomic data in Phase II.
Phase II
High-throughput sequencing of genomic clones was jointly performed by the MSCs. The JCVI produced an i3 assembly from the more than 19M trace files into contiguous stretches of genomic sequence called “contigs” and “scaffolds”. The JCVI and VectorBase are currently predicting Ixodes gene sets based on the i3 assembly (0.5 annotation) that will later be merged to produce a 1.1 gene set. Input from the research community will be essential to analyze and improve these automated gene predictions.
IGP data are uploaded at regular intervals to two public databases; the National Center for Biotechnology Information (GenBank) and VectorBase.
|




